Nanopore genetic sequencing is an innovative technique where the graphical interface software Rapid nanpOre Brain IntraoperatIve classificatioN (ROBIN) presents sequencing results of patients in real-time as data is being analysed and interpreted from a single assay (Deacon et al., 2025). The World Health Organisation (WHO) classification of tumours of the central nervous system (CNS) has placed emphasis on the routine use of molecular testing, as there are more than 100 types of brain tumours with defined mutations in the absence of defining morphological features (Louis et al., 2021; Sissons and Martin, 2025). This does not eliminate the use of immunohistochemistry and other microscopic analytical techniques; however, nanopore sequencing has provided its validation status through its effective sequencing, which surpasses other array technologies, its low price, and quick turnaround time. According to The Brain Tumour Charity, brain cancers are attributed to a high death rate of children and adults under the age of 40 (Sissons and Martin, 2025).
ROBIN was developed by researchers and clinicians at the University of Nottingham and Nottingham University Hospitals. This has significantly helped to improve time efficiency in diagnosis and surgical planning in patients with glioblastoma, an aggressive brain cancer (Sissons and Martin, 2025).
The standardised guidelines for brain cancer diagnosis are to extract a sample from the tumour during surgery, which is known as a biopsy. It undergoes a sequential examination under a microscope in a pathology laboratory, with a definitive diagnosis within eight weeks. This lowers the treatment options, as during the review consultation, the stage of the cancer has already grown. Surgeons aim to undergo radical surgery to remove the tumour if this has not been done in the initial operation. In some cases, patients undergo multiple surgeries to ensure all the tumour has been removed successfully (Sissons and Martin, 2025).
On the contrary, with the introduction of nanopore genetic sequencing, the time taken to diagnose the type of brain tumour transformed from weeks to 90 minutes with a 100% success rate. The researchers, Deacon et al. (2025), further added that the results received intraoperatively had a turnaround time of under two hours and a robust tumour classification within minutes. This is evidenced by 50 operations where neuropathologists were able to examine a small amount of genetic material, deoxyribonucleic acid (DNA), and select the parts to explore in detail for a faster result. The surgeons would receive a response while patients undergo radical surgery. It will allow other treatment modalities like radiation and chemotherapy after the operation. Moreover, there would be a decrease in the risk of anxiety in patients during the eight-week period for waiting for results (Sissons and Martin, 2025). This is a prodigious step, as with the comprehensive classification of brain tumours, it is essential to identify the type of tumour to provide the necessary treatment.
The success of this novel genetic technology is evidenced by patient X, who was diagnosed with a brain tumour in 2023 at age 1. The patient has undergone two brain surgeries in the past year, followed by a high dose of chemotherapy to lower the risk of relapse (cancer returning). The time taken from patient X’s surgery to identifying the type of tumour was two weeks, and the patient’s mother described it as ‘pure hell and anxiety.’
However, the mother was relieved when the nanopore sequencing technology was established, as it could have prevented doing the additional brain surgery at the time. The patient is currently in remission and is supported by the Young Lives vs Cancer charity (Sissons and Martin, 2025).
Another example is a paediatric patient who underwent subtotal resection surgery, while standard of care testing was performed. An additional operation was required, and it was identified to be a RELA::ZFTA mutation ependymoma 19 days after the surgery.
Ependymoma arises from ependymal cells that align the ventricles of the brain and the spinal cord canal. It is subdivided into three types that vary in location, genetic mutations, and DNA methylation profile (Park et al., 2023). Supratentorial ependymoma arises in the cerebral hemispheres of the brain. Posterior fossa ependymoma originates in the posterior fossa and cerebellum. Spinal ependymoma occurs in the spinal cord and is linked with neurofibromatosis type 2 (NF2) genetic mutation or deletion (Park et al., 2023).
Conversely, if nanopore-based analysis were applied, the additional surgery would be unnecessary. Furthermore, rare tumours are also diagnostically challenging and may delay the start of appropriate treatment (Deacon et al., 2025).
What Is Nanopore Genetic Sequencing?
Nanopore DNA sequencing helps to determine the specific sequence of nucleotides in the genetic material. There are four nitrogen-rich bases: Adenine (A), Thymine (T), Guanine (G), and Cytosine (C). Please see Figure 1. The DNA has two strands of polynucleotides, where one of these strands is signalled to read through tiny pores referred to as nanopores. This is found in the membrane (Wetterstrand, 2025). Different nucleotides of DNA are detected by the presence of changes in the electrical current. Please see Figure 2. This helps to identify complex sections in the human genome (Wetterstrand, 2025).
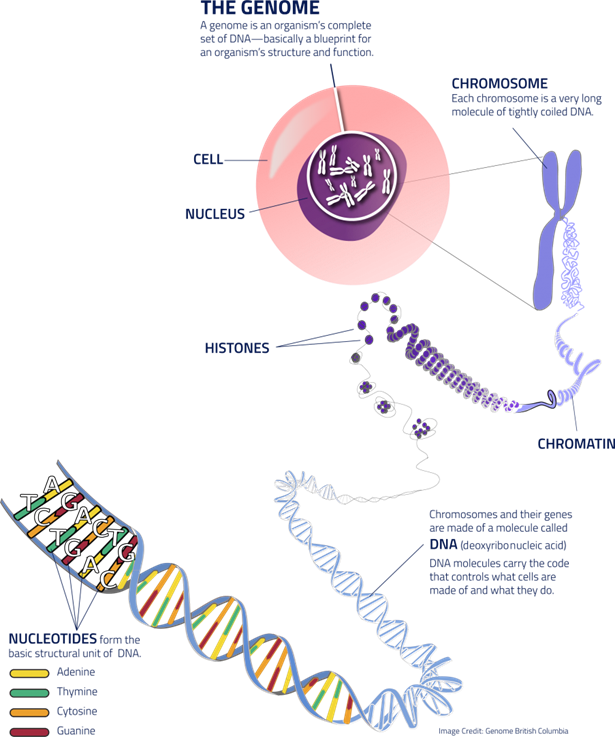
Figure 1: Structure of DNA
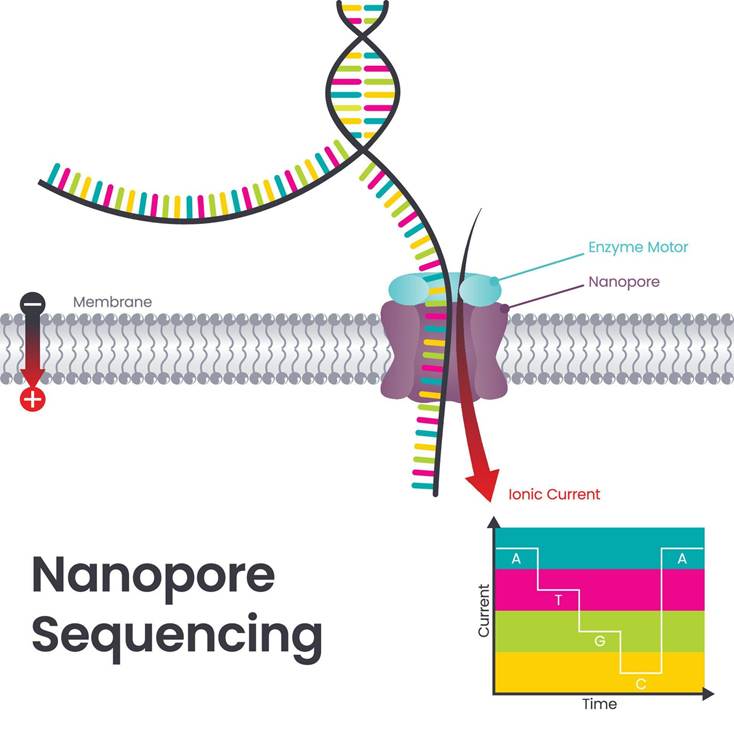
Figure 2 Nanopore Sequencing.
Three fresh tissue samples from the operating theatres were collected by Deacon et al. (2025). Each sample was between 5 and 25 mg of tissue, for further processing of DNA extraction and smear analysis intraoperatively. This helps to determine which would be suitable for sequencing with a higher tumour cell fraction and no extensive gross necrosis (cell death). There is a web interface where ROBIN operates, and the sequencing data output is saved on a computer. The neuropathologist can then make a diagnosis in real time based on the classification of the position and size of the data.
There were three types of methylation classifiers: Sturgeon, CrosssNN, and random forest (RF) that were used to evaluate their performance in sequencing the data. The purpose of methylation classifiers is to accurately classify brain tumours based on their methylation profiles. This helps healthcare professionals produce a personalised treatment plan after determining the diagnosis and predicting the tumour prognosis.
An additional advantage of methylation classifiers is overcoming histopathological challenges, where, in some patient cases, the possibility of obtaining ambiguous results may occur. Another limitation that may arise is a mismatch in information between histology and molecular profiling. There are two main types: not otherwise specified (NOS) cases or not elsewhere classified (NEC) cases (Park et al., 2023). Thus, methylation classifiers can facilitate decision-making.
On the contrary, while methylation patterns are insightful, it does not fully capture the heterogeneous nature of brain tumours. There is a strong likelihood of alternative epigenetic mechanisms, modification of histone proteins that stabilise the DNA, chromatin remodelling, and non-coding RNA regulation. Collectively, they formulate gene expression patterns in tumour cells (Bure et al., 2022).
What type of results were established intraoperatively via this technique?
There are several types of genetic mutations that can be identified by ROBIN: copy number variation (CNV), MGMT promoter methylation, single-nucleotide variation (SNV), and structural variants in the 50 patients analysed at Nottingham University Hospitals. This is achieved in real-time via native strong long-read sequencing. A complete diagnosis was achieved within 24 hours due to candidate fusions during the sequencing process (Deacon et al., 2025).
Single Nucleotide Variant
Single Nucleotide Variation (SNV) refers to a change within a single nucleotide in a DNA sequence that can occur in germ cells (sex cells) or somatic cells (body cells). As illustrated in Figure 3, three of the bases, cytosine (C), Adenine (A), and Thymine (T), produce the amino acid histidine. Amino acids are the building blocks that make protein. A change in the single nucleotide from A to C is considered an SNV. This produces a non-functioning protein.
In the research study by Deacon et al. (2025), several SNVs were discovered: the TERT promoter mutation was found in glioblastoma. TP53 mutation in astrocytoma. p.D463H (c.1387G > C) missense mutation in the PRKCA gene, classified as chordoid glioma, and was confirmed in 9 of 35 reads. A missense mutation is a type of SNV where there is a change in the amino acid, for instance, from histidine to arginine, but the protein product may be functional or non-functional. This illustrates that ROBIN is limited to high confidence when there are equally high variant allele frequencies (Deacon et al., 2025). However, in cases when there is a low tumour fraction, it can be resolved by additional panel testing.
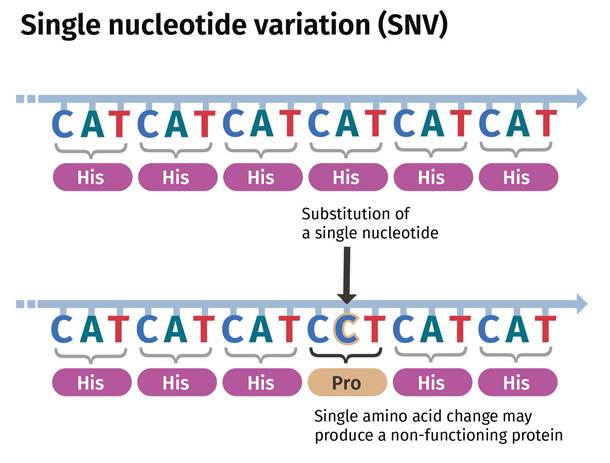
Figure 3: An example of a Single Nucleotide Variation (SNV)
Copy Number Variation
Copy Number Variation (CNV) is defined as sections of the genome repeated or copied, causing genetic diversity and susceptibility to disease. There are two common CNVs: deletion and duplication. Please see Figure 4.
According to Deacon et al.’s (2025) study, they revealed that most copy number profiles were associated with glioblastoma, where a gain of chromosome 7 and a loss of chromosome 10 were identified. Patients with oligodendroglioma were characterised with a loss of both chromosomes 1p and 19q. A patient with anaplastic oligodendroglioma classified as WHO Grade 3 had additional chromosome arms deleted (Deacon et al., 2025). Overall, the molecular genetic signatures of glioblastoma and oligodendrogliomas were identified.
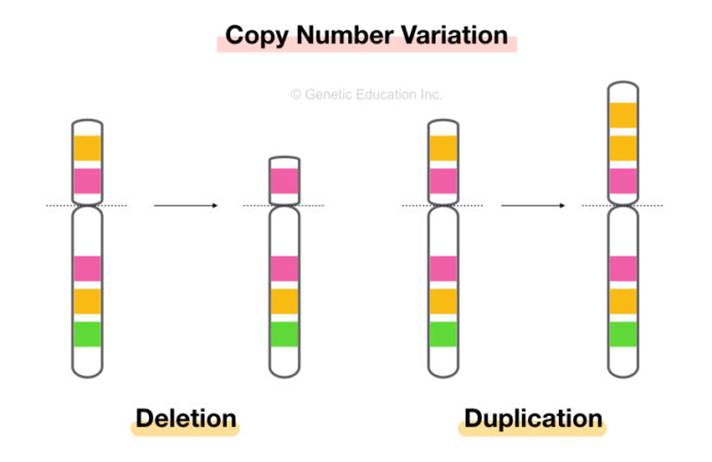
Figure 4: Copy Number Variation (CNV)
MGMT promoter methylation
DNA methylation of the amino acid cytosine occurs during cell divisions and can facilitate histogenesis. Methylation is the addition of the methyl group (CH3) to the molecule or the substitution of a methyl group. Histogenesis is defined as the growth and differentiation of cells into tissues and organs during embryonic development.
DNA methylation is an essential epigenetic modification that regulates gene expression and maintains genomic stability (Moore, Le, and Fan, 2012). Additional functions include genomic imprinting, X chromosome inactivation, and suppression of transposable elements (Park et al., 2023). Also known as transposons or jumping genes, transposable elements are mobile genetic sequences that can cause deletions, inversions, and other genomic changes.
The origin of cells during the neoplastic stage can be identified via nanopore sequencing technology based on the DNA methylation profiling status (Deacon et al., 2025). In the methylome analysis of patients with gliomas, clinical features, prognosis, and responses to treatment are distinctive between the subtypes (Reuss et al., 2015). By understanding how methylation influences the genome of cancer cells, target genes and signalling pathways that are dysregulated serve as potential therapeutic targets (Lewandowska et al., 2011). Hence, the WHO 2021 classification of tumours places emphasis on methylome testing.
O6-Methylguanine-DNA methyltransferase (MGMT) is a DNA repair enzyme that removes the methyl group at the O6 position of guanine added to DNA of glioma by the chemotherapeutic agent temozolomide and transfers the alkyl group to itself. Therefore, MGMT halts the mode of action of temozolomide, causing a neutralising effect(Sharma, 2024). Instead of methyl-guanine joining with cytosine, it joins with thymine, forming a mismatch during DNA replication (Hideaki et al., 2018).
As illustrated in Figure 5, there is a positive correlation between MGMT promoter methylation and MGMT protein expression. The MGMT protein cannot be produced if the MGMT is methylated because the genetic information cannot be copied (replication), transcribed, or translated to form a protein (Hideaki et al., 2018).
Moreover, this reaction occurs once per MGMT molecule. It is corrected by mismatch repair genes that remove the thymine, but again, the thymine is reinserted. This is now termed as a futile mismatch repair, where the tumour DNA becomes vulnerable, and the tumour cell undergoes apoptosis. However, if the tumour cell can regenerate new MGMT molecules, it can repair the damage caused by temozolomide (Sharma, 2024).
The promoter is the start of the DNA sequence. Methylation of the MGMT promoter decreases MGMT expression. Thus, MGMT promoter methylation is a biomarker to identify patients with low-grade IDH-mutant gliomas (80%) who respond positively to temozolomide (Hideaki et al., 2018; Sharma, 2024). Patients with IDH-mutant gliomas commonly express the Glioma CpG Island Methylator Phenotype (G-CIMP), with CpG hypermethylation in genomic regions. These patients have MGMT promoter methylation and are sensitive to temozolomide.
Furthermore, patients with WHO grade 3 and 4 malignant gliomas (astrocytoma and glioblastoma) are also sensitive towards Temozolomide. MGMT promoter methylation is found in 35 to 45% of tumours (Hideaki et al., 2018; Sharma, 2024). This indicates that low-grade gliomas have higher MGMT promoter methylation compared to high-grade gliomas.
Deacon et al. (2025) illustrate that 44 of 46 in the prospective cohort (forward in time or data) had MGMT status. 25 of 29 had a retrospective cohort (past events in time or data). There was one patient with a methylation score near the cutoff value. This could be associated with systematic approaches between the array and nanopore thresholds. Thus, a low level of methylation of MGMT may still be sensitive to temozolomide (Deacon et al., 2025).
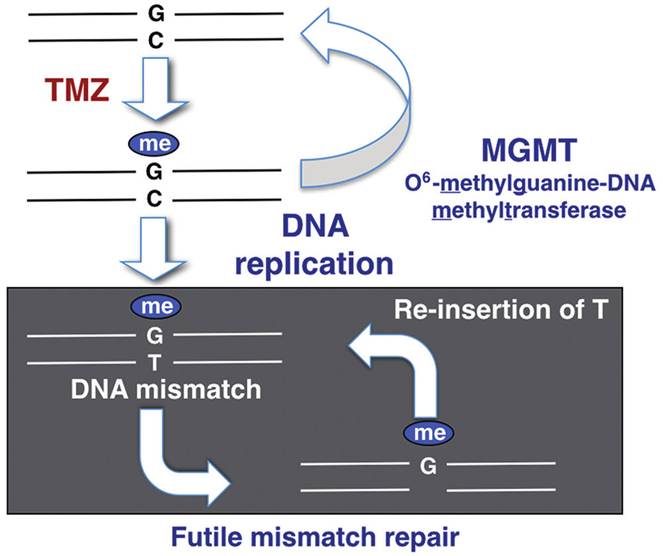
Figure 5: MGMT methylation
Structural variants
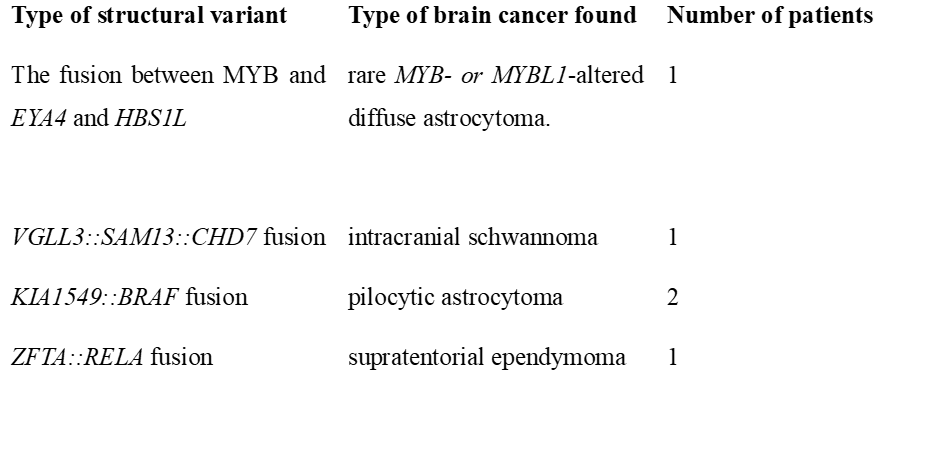
Through Nanopore sequencing, five cases had relevant fusions; please see Table 1. Most glioblastomas did not have fusions, except for one that had SEPTIN14::EGFR fusion. In addition, it is common that MYB- or MYB1L, with PCDHGA1, MMP16, and MAML2, are formed in rare MYB- or MYBL1-altered diffuse astrocytoma. However, Deacon et al. (2025) discovered a fusion between MYB and alternative partner genes. Please see Table 1 for additional structural variants.
Table 1: Structural variants found in brain cancer patients

The advantages and disadvantages of Nanopore
There are positive and negative applications of nanopore sequencing. On one hand, it is time-efficient and helps prevent additional surgeries beyond the first attempt. However, there is a challenge in forming a balance intraoperatively where they rapidly sample, process, and create DNA of good yield and read length to detect additional molecular features (Deacon et al., 2025). It relies on ultra-low read depth, where information focused on binary methylation can be obtained, and most sites and specialised tools are needed for accuracy. They are not suitable for identifying additional data, single-nucleotide variation, and structural variants. This is primarily due to high confidence and low coverage, where additional RNA and DNA sequencing is necessary.
Moreover, the turnaround time was less than two hours. 38% (76%) of cases were prospective and classified using two or more classifiers within 1 hour of sequencing, which vary in performance. Sturgeon and CrosssNN were able to classify the data within minutes, whereas random forest (RF) was slow. Initially, the poor performance of classification was associated with a short supply of data and low coverage of CpG sites. Sturgeon overconfidently misclassified and switched class mid-sequencing. This highlights that there are advantages and disadvantages in every analytical technique, with the aim of ensuring the benefits overcome the risks.
There are other substantial challenges that arise in methylome testing. It cannot classify all brain tumours and may produce matches in error with high scores. The cause of these limitations is the inability to access all reference methylation data profiles for all types of brain tumours. This includes novel types that have been identified and are not readily accessible in open databases (Park et al., 2023).
On a global level, not all hospitals are equipped with specialised laboratory techniques nor have medical insurance to cover methylation studies because the cost of the apparatus and its reagents for maintenance in Asian countries is estimated to be three times higher than in Western countries (Park et al., 2023).
Overall, nanopore sequencing has significant advantages in comparison to other genetic and array technologies. It has motivated interest in multiple hospitals across the UK to establish their own nanopore genetic testing section in their laboratories. Alternatively, on a global level, an increased need for equity and accessible resources for the developed and underdeveloped world for methylation studies is a limitation in the global mission to improve cancer diagnosis and treatment and meet the WHO’s sustainable development goals (SDGs) by 2030 (Sissons and Martin, 2025; Park et al., 2024).
References
Abe, H., Natsumeda, M., Kanemaru, Y., Watanabe, J., Tsukamoto, Y., Okada, M., Yoshimura, J., Oishi, M. And Fujii, Y. (2018). Mgmt Expression Contributes To Temozolomide Resistance In H3k27m-Mutant Diffuse Midline Gliomas And Mgmt Silencing To Temozolomide Sensitivity In Idh-Mutant Gliomas. Neurologia Medico-Chirurgica, 58(7), Pp.290–295. Doi:Https://Doi.Org/10.2176/Nmc.Ra.2018-0044.
Bure, I.V., Nemtsova, M.V. and Kuznetsova, E.B. (2022). Histone Modifications and Non-Coding RNAs: Mutual Epigenetic Regulation and Role in Pathogenesis. International Journal of Molecular Sciences, [online] 23(10), p.5801. doi:https://doi.org/10.3390/ijms23105801.
Deacon, S., Cahyani, I., Holmes, N., Fox, G., Munro, R., Wibowo, S., Murray, T., Mason, H., Housley, M., Martin, D., Sharif, A., Patel, A., Goldspring, R., Brandner, S., Sahm, F., Smith, S., Paine, S. and Loose, M. (2025). ROBIN: A unified nanopore-based assay integrating intraoperative methylome classification and next-day comprehensive profiling for ultra-rapid tumor diagnosis. Neuro-Oncology. [online] doi:https://doi.org/10.1093/neuonc/noaf103.
Jin Woo Park, Lee, K., Eric Eunshik Kim, Kim, S.-I. and Park, S.-H. (2023). Brain Tumor Classification by Methylation Profile. Journal of Korean Medical Science, 38(43). doi:https://doi.org/10.3346/jkms.2023.38.e356.
Lewandowska, J. and Bartoszek, A. (2011). DNA methylation in cancer development, diagnosis and therapy–multiple opportunities for genotoxic agents to act as methylome disruptors or remediators. Mutagenesis, 26(4), pp.475–487. doi:https://doi.org/10.1093/mutage/ger019.
Louis, D.N., Perry, A., Wesseling, P., Brat, D.J., Cree, I.A., Figarella-Branger, D., Hawkins, C., Ng, H.K., Pfister, S.M., Reifenberger, G., Soffietti, R., von Deimling, A. and Ellison, D.W. (2021). The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro-Oncology, 23(8). doi:https://doi.org/10.1093/neuonc/noab106.
Moore, L.D., Le, T. and Fan, G. (2012). DNA Methylation and Its Basic Function. Neuropsychopharmacology, [online] 38(1), pp.23–38. doi:https://doi.org/10.1038/npp.2012.112.
Reuss, D.E., Kratz, A., Sahm, F., Capper, D., Schrimpf, D., Koelsche, C., Hovestadt, V., Bewerunge-Hudler, M., Jones, D.T.W., Schittenhelm, J., Mittelbronn, M., Rushing, E., Simon, M., Westphal, M., Unterberg, A., Platten, M., Paulus, W., Reifenberger, G., Tonn, J.-C. and Aldape, K. (2015). Adult IDH wild type astrocytomas biologically and clinically resolve into other tumor entities. Acta Neuropathologica, [online] 130(3), pp.407–417. doi:https://doi.org/10.1007/s00401-015-1454-8.
Sharma, R. (2024) Methylguanine-DNA methyltransferase (MGMT) Available at: https://radiopaedia.org/articles/methylguanine-dna-methyltransferase-mgmt (Accessed: 14th November 2025)
Sissons, R. and Martin, D. (2025) New brain tumour test a game changer, experts say. Available at: https://www.bbc.co.uk/news/articles/cpvkmkk4pv4 (Accessed: 7th November 2025)
Wetterstrand, K. (2025) Nanopore DNA Sequencing Available at: https://www.genome.gov/genetics-glossary/Nanopore-DNA-Sequencing (Accessed: 13th November 2025)
Leave a comment